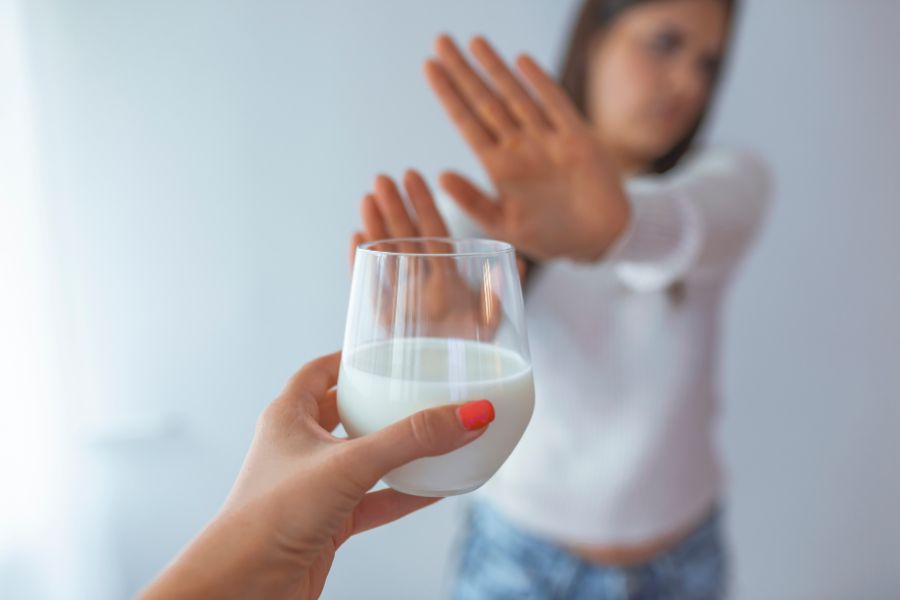
While some people wonder, “what is whole milk?” and if it’s good for them, others can’t digest any form of dairy. The discomfort you experience after indulging in a glass of warm milk or a bowl of ice cream results from lactose intolerance. Guess what? The condition is way more common than you think.
Up to 75% of the world’s population have some degree of lactose intolerance, meaning they lack the enzyme needed to digest lactose…
What Are Entry Requirements to Study Medicine?
Are you aspiring to follow a career path in medicine after completing your secondary education? Are you wondering about specific medical requirements? At least you know you must attain a high GPA or final high school score and good MCAT prep books to qualify for Medical Colleges.
Below we are going to focus on the different prerequisites to joining Medical School:
What High School Subjects Are Needed to Pursue
Any medical professional should have strong grades in mathematics and science subjects (Chemistry, Biology, and Physics).
For US Medical Schools, Mid-level mathematics with chemistry and any other …
Tips for People With ADHD to Try: 5 Techniques That Helps
Living with ADHD can be very difficult. The simplest things become frustrating because of the inability to concentrate or focus on many things simultaneously. This, in turn, leads to a decrease in productivity. When you don’t have access to some, you can use these techniques to avoid distractions, work efficiently and be happier.
5 Simple Techniques to Help You Cope with ADHD
1. Create a Routine
A research study on ADHD shows that one of the biggest challenges for people with ADHD is concentrating on different …
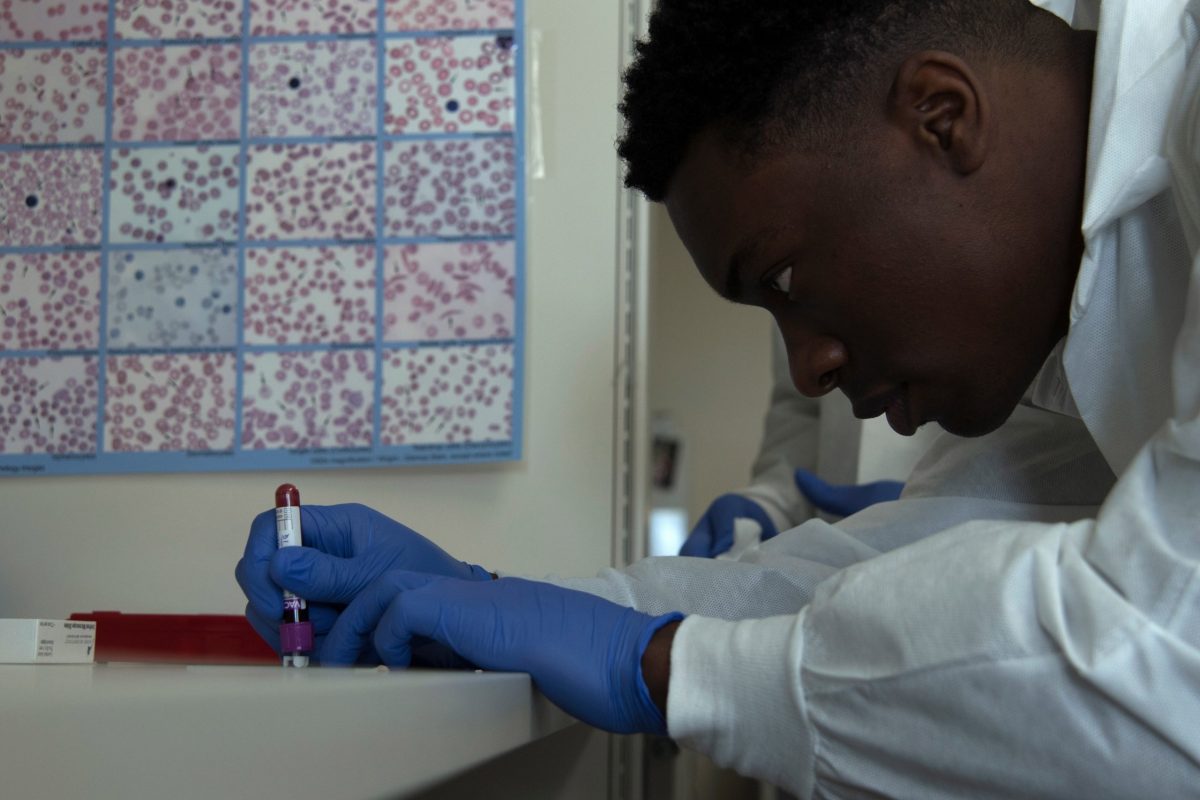
Prevention of Hospital Acquired Infection During COVID-19: a New Discovery Available
Early detection and prevention of hospital-acquired infections through passive monitoring devices help in the reduction of cases. COVID-19 is another threat to patients and health care workers in hospitals. Due to the inflation of COVID-19 cases, people avoid entering hospitals, specifically the emergency rooms, where many cases come in each day.
Moreover, the public is aware of the associated risks when exposed to infected people with COVID-19. A recent study developed a new tool to detect hospital-acquired infections with an association of five pathogens.
Hospital-acquired diseases monitoring
The investigators …
Atopic Dermatitis in Children: Skin Microflora Study
Atopic Dermatitis is a chronic skin condition and most common among children. The research study aims to identify how fungi and bacteria cause this skin condition. Moreover, it expects to determine basic measures to create a more advanced and effective future treatment. Healthy children and those with atopic dermatitis are welcome to join.
Commonly Asked Questions
What is included?
For Patients with Atopic Dermatitis (Eczema):
The child’s skin sample is needed at separate time points such as baseline, flare, and post flare. Visit scheduling and timing are necessary to conduct this study.
Healthy …
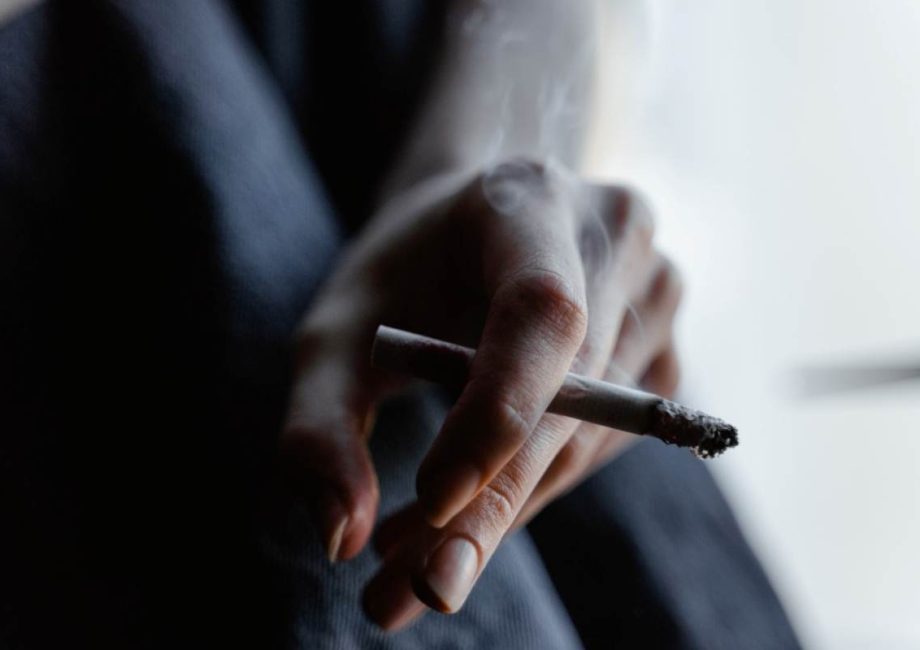
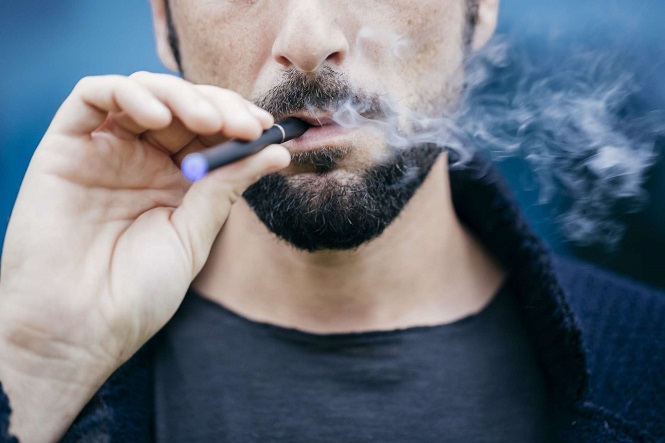
Vaping Doesn’t Differ From Smoking: It Can Cause the Same Chemical Changes to Our Genome
Smoking can harm your heart, lungs, and most of your organs. People who usually want to live a healthier lifestyle decide to kick smoking out of their habits and become less dependent on it. However, people who smoke when they want to quit smoking are usually tempted to turn to vaping devices.
As part of their transition from cigarette smoking to none at all, smokers consider that using vaping devices or e-cigarettes can help them adjust eventually. Although it is less harmful than smoking nicotine, vaping, according to scientists, is still unsafe says Industry leaders.
Vaping Causes …
Genomics Education Week for Healthcare Provider
The future of medicine changes because of discoveries such as genomics. Genomics plays a significant role in patient care and wellness. However, most healthcare providers are not aware and well equipped with knowledge on genomics. Raising awareness on genomics and the resources available will be covered in this event. Moreover, a social media campaign will be conducted during the educational week that primarily focuses on imparting genomic education. The activities included are webinars, social media such as Twitter and other platforms, panel discussion, and question and answer sessions.
The event details are as follows:
…
Genomics Branch: Computational and Statistical
The genomic and genetic analysis takes place through an intensive approach application which is the main focal point of the Computational and Statistical Genomics Branch.
Overview
The investigators of this branch are experts in genetic epidemiology and statistical genetics. These fields include mathematics, epidemiology, statistics, computer science, and molecular genetics for identifying disease susceptibility.
Moreover, experts’ objective is to expand knowledge on the complexity of diseases and advance new statistical approaches to evaluate and analyze the spectrum of genetic data. Computational biologists …